Lab: Multilayer networks
Last update: 03 March, 2026
#devtools::install_github("manlius/muxViz")
# library(muxViz)
library(emln)
library(infomapecology)
library(igraph)
library(bipartite)
library(tidyverse)
library(magrittr)
library(readxl)
library(ggalluvial)
library(ggplot2)
library(tidyr)
library(dplyr)
library(purrr)
library(network)
library(ggnetwork)In this exercise we will rely heavily on the EMLN package. See the full description and code in https://ecological-complexity-lab.github.io/emln_package/.
Code
1. Data structures: Monolayer networks
To understand multilayer network data structures we must first understand the monolayer data structures. We did this in our first exercise. Therefore, we assume you already know this!
The EMLN package has built in functions to convert between edge lists and matrices. You should follow the code here and make sure you understand it before delvoing into multilayer networks.
2. Data structures: Multilayer networks
We will again largely follow the guide of the EMLN package here. But before we do, let’s learn the data structures.
Multilayer network components
We can divide multilayer networks into those with and without interlayer edges:
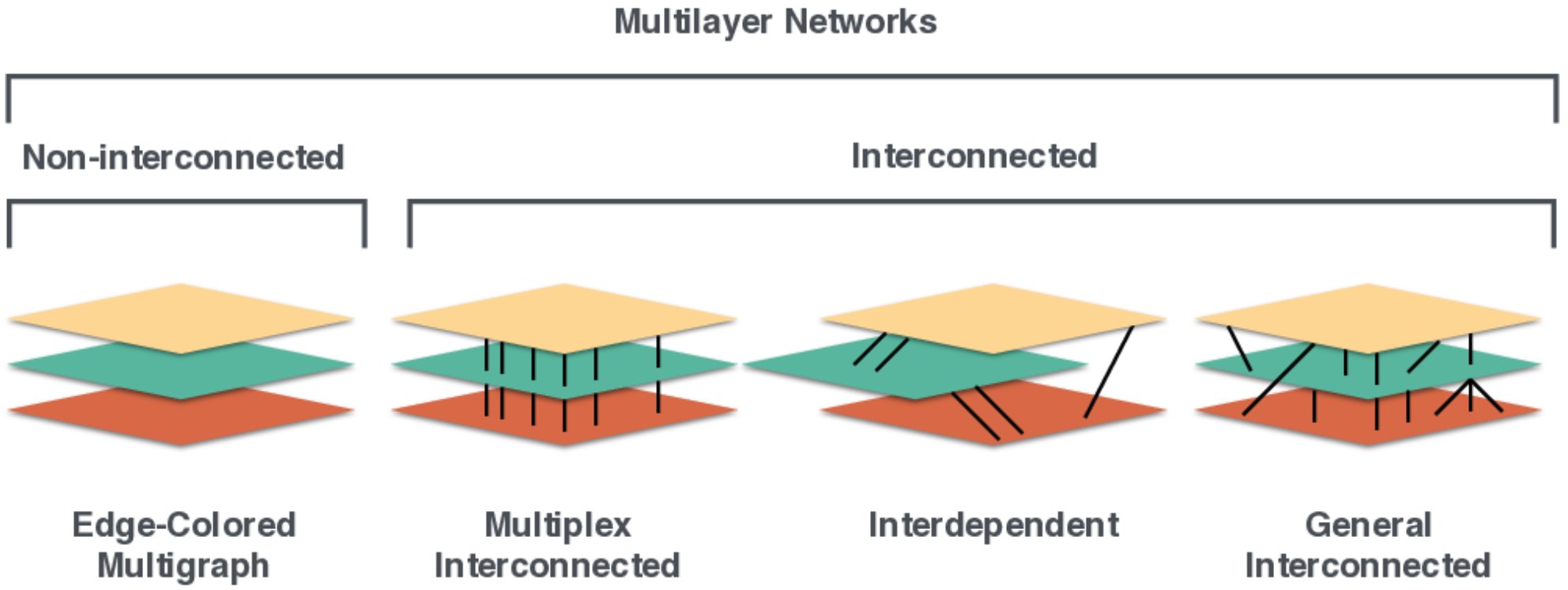
And having the following components:
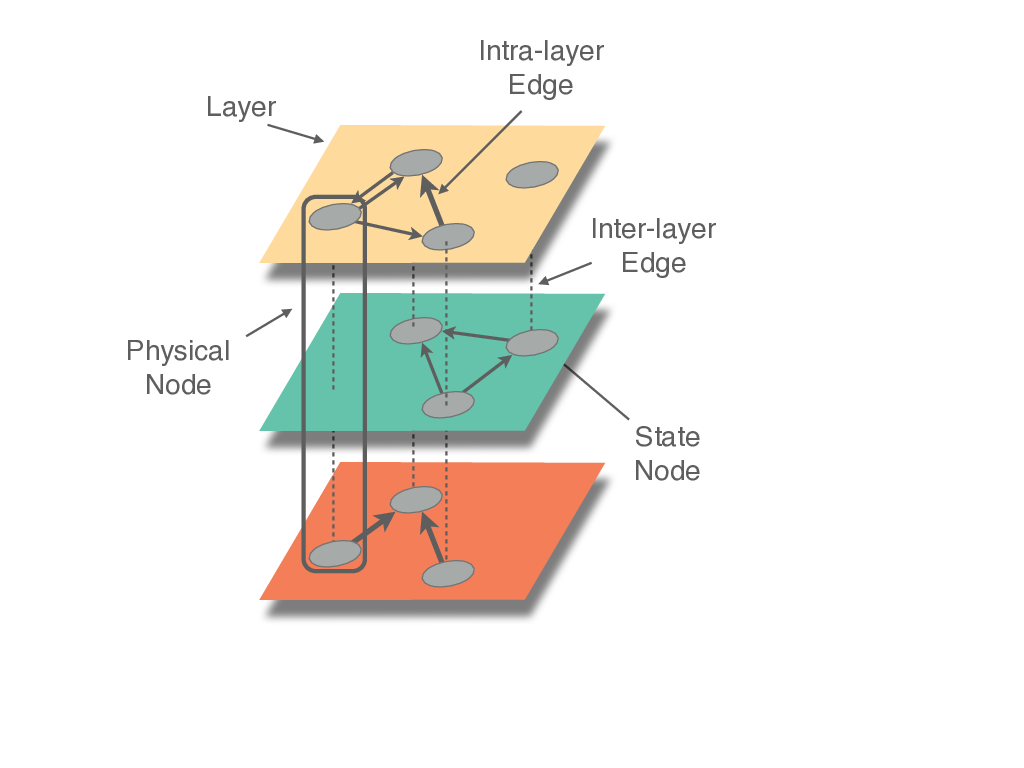
(Figures copy-righted to Manlio de Domenico, https://github.com/manlius/muxViz/blob/master/gui-old/theory/README.md)
Three representations of a multilayer network
We can represent a multilayer using a graph, an extended edge list or a tensor (matrix with more than 2 dimensions). Tensors can be flattened to supra-adjacency matrices. This can be seen in the middle panel.
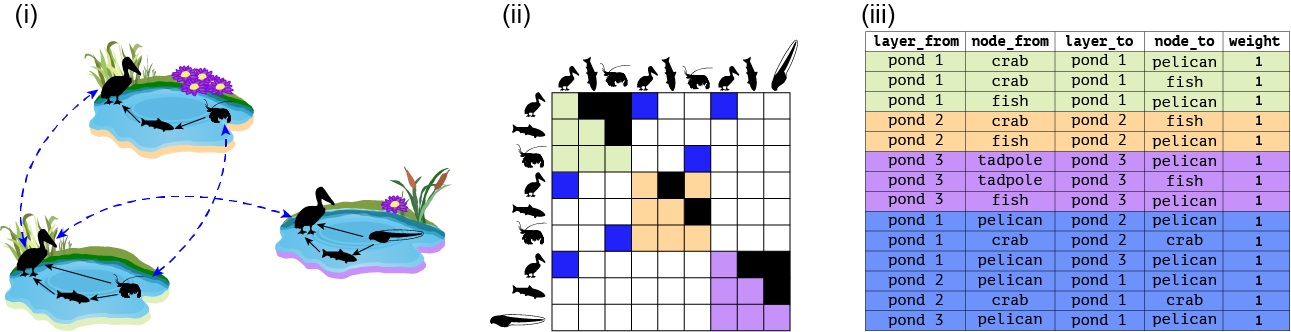
And here:
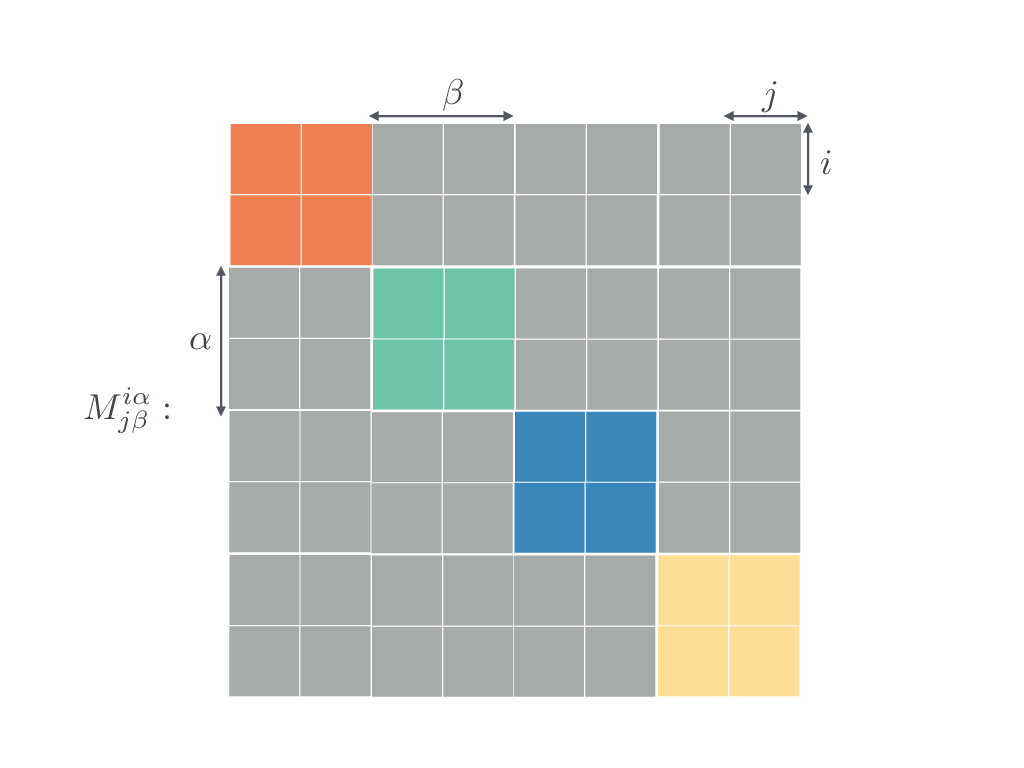
No interlayer links.
Now that we understand structures, let’s see how to encode them.
Toy example
This is the easiest case of disconnected layers. Here is the pond example without interlayer links:
pond_1 <- matrix(c(0,1,1,0,0,1,0,0,0), byrow = T, nrow = 3, ncol = 3, dimnames = list(c('pelican','fish','crab'),c('pelican','fish','crab')))
pond_2 <- matrix(c(0,1,0,0,0,1,0,0,0), byrow = T, nrow = 3, ncol = 3, dimnames = list(c('pelican','fish','crab'),c('pelican','fish','crab')))
pond_3 <- matrix(c(0,1,1,0,0,1,0,0,0), byrow = T, nrow = 3, ncol = 3, dimnames = list(c('pelican','fish','tadpole'),c('pelican','fish','tadpole')))
layer_attrib <- tibble(layer_id=1:3,
layer_name=c('pond_1','pond_2','pond_3'),
location=c('valley','mid-elevation','uphill'))
multilayer <- create_multilayer_network(list_of_layers = list(pond_1, pond_2, pond_3),
bipartite = F,
directed = T)## [1] "Layer attributes not provided, I added them (see layer_attributes in the final object)"
## [1] "Layer #1 processing."
## [1] "Done."
## [1] "Layer #2 processing."
## [1] "Done."
## [1] "Layer #3 processing."
## [1] "Done."
## [1] "Creating extended link list with node IDs"
## [1] "Organizing state nodes"## # A tibble: 8 × 5
## layer_from node_from layer_to node_to weight
## <chr> <chr> <chr> <chr> <dbl>
## 1 layer_1 fish layer_1 pelican 1
## 2 layer_1 crab layer_1 pelican 1
## 3 layer_1 crab layer_1 fish 1
## 4 layer_2 fish layer_2 pelican 1
## 5 layer_2 crab layer_2 fish 1
## 6 layer_3 fish layer_3 pelican 1
## 7 layer_3 tadpole layer_3 pelican 1
## 8 layer_3 tadpole layer_3 fish 1The multilayer class
Go over the resulting multilayer
class and see what it includes.
Real data
Let’s take an example with real data, using the data of Kefi et al 2016. This is a node-aligned multiplex network with three layers: trophic interactions, non-trophic positive interactions and non-trophic negative interactions. First, we will see how we can use matrices as input. Along the way we perform sanity checks such as making sure the number and identity of of species is the same in the 3 layers.
# Load and organize the trophic interactions matrix
chilean_TI <- read_delim('data/chilean_TI.txt', delim = '\t')
nodes <- chilean_TI[,2]
names(nodes) <- 'Species'
chilean_TI <- data.matrix(chilean_TI[,3:ncol(chilean_TI)])
dim(chilean_TI)## [1] 106 106dimnames(chilean_TI) <- list(nodes$Species,nodes$Species)
# Load and organize the negative non-trophic interactions matrix
chilean_NTIneg <- read_delim('data/chilean_NTIneg.txt', delim = '\t')
nodes <- chilean_NTIneg[,2]
names(nodes) <- 'Species'
chilean_NTIneg <- data.matrix(chilean_NTIneg[,3:ncol(chilean_NTIneg)])
dim(chilean_NTIneg)## [1] 106 106dimnames(chilean_NTIneg) <- list(nodes$Species,nodes$Species)
# Are all row and column names the same?
setequal(rownames(chilean_TI), rownames(chilean_NTIneg))## [1] TRUE## [1] TRUEchilean_NTIneg <- chilean_NTIneg[rownames(chilean_TI),colnames(chilean_TI)]
all(rownames(chilean_TI)==rownames(chilean_NTIneg))## [1] TRUE# Load and organize the negative non-trophic positive matrix
chilean_NTIpos <- read_delim('data/chilean_NTIpos.txt', delim = '\t')
nodes <- chilean_NTIpos[,2]
names(nodes) <- 'Species'
chilean_NTIpos <- data.matrix(chilean_NTIpos[,3:ncol(chilean_NTIpos)])
dim(chilean_NTIpos)## [1] 106 106dimnames(chilean_NTIpos) <- list(nodes$Species,nodes$Species)
# Are all row and column names the same?
setequal(rownames(chilean_TI), rownames(chilean_NTIpos))## [1] TRUE## [1] TRUEchilean_NTIpos <- chilean_NTIpos[rownames(chilean_TI),colnames(chilean_TI)]
all(rownames(chilean_TI)==rownames(chilean_NTIpos))## [1] TRUE## [1] 4623# Create the multilayer object
layer_attributes <- tibble(layer_id=1:3, layer_name=c('TI','NTIneg','NTIpos'))
multilayer_kefi <- create_multilayer_network(list_of_layers = list(chilean_TI,chilean_NTIneg,chilean_NTIpos),
interlayer_links = NULL,
layer_attributes = layer_attributes,
bipartite = F,
directed = T)## [1] "Layer #1 processing."
## [1] "Done."
## [1] "Layer #2 processing."
## [1] "Some nodes have no interactions. They will appear in the node table but not in the edge list"
## [1] "Done."
## [1] "Layer #3 processing."
## [1] "Some nodes have no interactions. They will appear in the node table but not in the edge list"
## [1] "Done."
## [1] "Creating extended link list with node IDs"
## [1] "Organizing state nodes"Now, we can try and use link lists as input. We will convert the matrices to link lists first.
# This is for the Kefi data set
chilean_TI_ll <- matrix_to_list_unipartite(x = chilean_TI, directed = TRUE)$edge_list
chilean_NTIneg_ll <- matrix_to_list_unipartite(x = chilean_NTIneg, directed = TRUE)$edge_list## [1] "Some nodes have no interactions. They will appear in the node table but not in the edge list"## [1] "Some nodes have no interactions. They will appear in the node table but not in the edge list"multilayer_kefi_ll <- create_multilayer_network(list_of_layers = list(chilean_TI_ll,chilean_NTIneg_ll,chilean_NTIpos_ll), layer_attributes = layer_attributes, bipartite = F, directed = T)## [1] "Layer #1 processing."
## [1] "Input: an unipartite edge list"## [1] "Done."
## [1] "Layer #2 processing."
## [1] "Input: an unipartite edge list"## [1] "Done."
## [1] "Layer #3 processing."
## [1] "Input: an unipartite edge list"## [1] "Done."
## [1] "Creating extended link list with node IDs"
## [1] "Organizing state nodes"# Check that the set of links is the same in both
links_mat <- multilayer_kefi$extended_ids %>% unite(layer_from, node_from, layer_to, node_to)
links_ll <- multilayer_kefi_ll$extended_ids %>% unite(layer_from, node_from, layer_to, node_to)
any(duplicated(links_mat$layer_from))## [1] FALSE## [1] FALSE## [1] TRUEAdding interlayer links
Toy example
In multilayer networks we have additional information we need to incorporate about the interlayer edges. Let’s see first the pond example.
# Create the ELL tibble with interlayer links.
interlayer <- tibble(layer_from=c('pond_1','pond_1','pond_1'),
node_from=c('pelican','crab','pelican'),
layer_to=c('pond_2','pond_2','pond_3'),
node_to=c('pelican','crab','pelican'),
weight=1)
# This is a directed network so the links should go both ways, even though they are symmetric.
interlayer_2 <- interlayer[,c(3,4,1,2,5)]
names(interlayer_2) <- names(interlayer)
interlayer <- rbind(interlayer, interlayer_2)
# An example with layer attributes and interlayer links
multilayer_unip_toy <- create_multilayer_network(list_of_layers = list(pond_1, pond_2, pond_3),
interlayer_links = interlayer,
layer_attributes = layer_attrib,
bipartite = F,
directed = T)## [1] "Layer #1 processing."
## [1] "Done."
## [1] "Layer #2 processing."
## [1] "Done."
## [1] "Layer #3 processing."
## [1] "Done."
## [1] "Creating extended link list with node IDs"
## [1] "Organizing state nodes"## # A tibble: 14 × 5
## layer_from node_from layer_to node_to weight
## <chr> <chr> <chr> <chr> <dbl>
## 1 pond_1 fish pond_1 pelican 1
## 2 pond_1 crab pond_1 pelican 1
## 3 pond_1 crab pond_1 fish 1
## 4 pond_2 fish pond_2 pelican 1
## 5 pond_2 crab pond_2 fish 1
## 6 pond_3 fish pond_3 pelican 1
## 7 pond_3 tadpole pond_3 pelican 1
## 8 pond_3 tadpole pond_3 fish 1
## 9 pond_1 pelican pond_2 pelican 1
## 10 pond_1 crab pond_2 crab 1
## 11 pond_1 pelican pond_3 pelican 1
## 12 pond_2 pelican pond_1 pelican 1
## 13 pond_2 crab pond_1 crab 1
## 14 pond_3 pelican pond_1 pelican 1Real example with a bipartite network
In this example we will work first with intralayer edges, and then add interlayer edges. We will use the spatial data of Gilarranz et al 2014, J Anim Ecol. The data contains 12 layers (patches) of plant-pollinator interactions. We use the original data, provided in Excel to show how working with real data feels like. This illustrates the workflow discussed in the paper.
Let’s start with intralayer edges (disconnected set of layers)
# Import layers
Sierras_matrices <- NULL
for (layer in 1:12){
d <- suppressMessages(read_excel('data/Gilarranz2014_Datos Sierras.xlsx', sheet = layer+2))
web <- data.matrix(d[,2:ncol(d)])
rownames(web) <- as.data.frame(d)[,1]
web[is.na(web)] <- 0
Sierras_matrices[[layer]] <- web
}
names(Sierras_matrices) <- excel_sheets('data/Gilarranz2014_Datos Sierras.xlsx')[3:14]
# Layer dimensions
sapply(Sierras_matrices, dim)## Amarante Barrosa Cinco Cerros Difuntito Difuntos El Morro La Brava La Chata La Paja Piedra Alta Vigilancia Volcan
## [1,] 39 36 33 38 25 17 24 33 28 22 34 34
## [2,] 67 58 63 79 69 67 58 61 48 66 74 68Why do the dimensions of the matrices matter?
Use EMLN to create the multilayer object:
layer_attrib <- tibble(layer_id=1:12, layer_name=excel_sheets('data/Gilarranz2014_Datos Sierras.xlsx')[3:14])
multilayer_sierras <- create_multilayer_network(list_of_layers = Sierras_matrices, bipartite = T, directed = F, layer_attributes = layer_attrib)See the extended edge list:
## # A tibble: 2,238 × 5
## layer_from node_from layer_to node_to weight
## <chr> <chr> <chr> <chr> <dbl>
## 1 Amarante Anthomyiidae Amarante Achyrocline satureoides 1
## 2 Amarante Apis mellifera Amarante Achyrocline satureoides 1
## 3 Amarante Augochlorella ephyra Amarante Achyrocline satureoides 1
## 4 Amarante Camponotus sp Amarante Achyrocline satureoides 1
## 5 Amarante Chilicola sp Amarante Achyrocline satureoides 1
## 6 Amarante Mischocyttarus drewseni Amarante Achyrocline satureoides 1
## 7 Amarante Parachytas sp Amarante Achyrocline satureoides 1
## 8 Amarante Sarcophagidae Amarante Achyrocline satureoides 1
## 9 Amarante Villa sp Amarante Achyrocline satureoides 1
## 10 Amarante Augochlora semiramis Amarante Alophia lahue 1
## # ℹ 2,228 more rowsTo illustrate working with interlayer links, we will connect a pollinator to itself between layers, and weigh these links by the distance between the layers. We emphasize that this is for illustration purposes only; there is no hypothesis behind this exercise. We already calculated the distances between the 12 patches. We can also do that for plants, but we keep this simple for the sake of the example.
dist <- data.matrix(read_csv('data/Gilarranz2014_distances.csv'))
dimnames(dist) <- list(layer_attrib$layer_name, layer_attrib$layer_name)
# Get the dispersal matrix, for pollinators only. This matrix shows which pollinator is in which layer
dispersal <- multilayer_sierras$extended %>% group_by(node_from) %>% dplyr::select(layer_from) %>% table()
dispersal <- 1*(dispersal>0)
Sierras_interlayer <- NULL
for (s in rownames(dispersal)){ # For each pollinator
x <- dispersal[s,]
locations <- names(which(x!=0)) # locations where pollinator occurs
if (length(locations)<2){next}
pairwise <- combn(locations, 2)
# Create interlayer edges between pairwise combinations of locations
for (i in 1:ncol(pairwise)){
a <- pairwise[1,i]
b <- pairwise[2,i]
weight <- dist[a,b] # Get the interlayer edge weight
Sierras_interlayer %<>% bind_rows(tibble(layer_from=a, node_from=s, layer_to=b, node_to=s, weight=weight))
}
}
# Re-create the multilayer object with interlayer links
multilayer_sierras_interlayer <- create_multilayer_network(list_of_layers = Sierras_matrices, bipartite = T, directed = F, layer_attributes = layer_attrib, interlayer_links = Sierras_interlayer)
# See the interlayer links
multilayer_sierras_interlayer$extended %>% filter(layer_from!=layer_to)
multilayer_sierras_interlayer$extended_ids %>% filter(layer_from!=layer_to)Plot the distribution of intralayer and interlayer links
Converting to supra-adjacency matrices and igraph
For some analysis it is easier to use SAM or iograph objects than an
extended edge list. It depends on how the algorithm/function works. See
how to convert a multilayer object in the EMLN
guidelines.
Exercise: multilayer data structures
Choose a temporal data set from those included in EMLN package and build a multilayer network. Draw interlayer edges by connecting the same species across layers (from t to t+1).
3. Plotting
We will plot a simulated multilayer network using ggnetwork as we learned in the previous classes. This plotting code is easily adjustable to empirical networks.
# 1. "Standing" Projection Function
# x_data -> depth (skewed)
# y_data -> height (preserved)
# offset -> shift along the 'depth' axis
project_standing <- function(x, y, offset) {
# We use a 45-degree skew for depth.
# X is skewed by Y slightly to give perspective, then shifted by offset.
list(x = (x * 0.5) + offset, y = y + (x * 0.2))
}
# Simulated layers
set.seed(42)
offsets <- c(0, 1.5, 3) # The "Z" distance between plates
layer_names <- c("Time 1", "Time 2", "Time 3")
n_nodes <- 50
all_edges <- data.frame()
plane_data <- data.frame()
node_coords_list <- list()
# 2. Generate and Project "Standing" Layers
for (i in 1:3) {
adj <- matrix(sample(0:1, n_nodes^2, replace = TRUE, prob = c(0.96, 0.04)), n_nodes, n_nodes)
net <- network(adj, directed = FALSE)
# Using 'fruchtermanreingold' inside the "plates" for a more organic look
# since circle layout can look a bit rigid when tilted this way.
df <- ggnetwork(net, layout = "fruchtermanreingold")
# Apply the standing projection
proj_start <- project_standing(df$x, df$y, offsets[i])
proj_end <- project_standing(df$xend, df$yend, offsets[i])
df$x <- proj_start$x; df$y <- proj_start$y
df$xend <- proj_end$x; df$yend <- proj_end$y
df$layer <- layer_names[i]
all_edges <- rbind(all_edges, df)
node_coords_list[[i]] <- df %>% select(x, y, vertex.names) %>% distinct()
# Standing Plane Background (The "Plate")
corners <- data.frame(px = c(0, 0, 1, 1), py = c(0, 1, 1, 0))
proj_c <- project_standing(corners$px, corners$py, offsets[i])
plane_data <- rbind(plane_data, data.frame(x = proj_c$x, y = proj_c$y, layer = layer_names[i]))
}
# 3. Generate 30 Random Inter-layer Links
interlink_indices <- data.frame(
from_node = sample(1:n_nodes, 30, replace = TRUE),
to_node = sample(1:n_nodes, 30, replace = TRUE),
connection = c(rep("1-2", 15), rep("2-3", 15))
)
interlayer_segments <- data.frame()
for (j in 1:nrow(interlink_indices)) {
idx <- interlink_indices[j, ]
l1 <- if(idx$connection == "1-2") 1 else 2
l2 <- if(idx$connection == "1-2") 2 else 3
interlayer_segments <- rbind(interlayer_segments, data.frame(
x = node_coords_list[[l1]]$x[idx$from_node],
y = node_coords_list[[l1]]$y[idx$from_node],
xend = node_coords_list[[l2]]$x[idx$to_node],
yend = node_coords_list[[l2]]$y[idx$to_node]
))
}
# 4. Final Visualization
ggplot() +
# Background Planes (Standing Plates)
geom_polygon(data = plane_data, aes(x = x, y = y, group = layer),
fill = "white", alpha = 0.3, color = "grey70") +
# INTER-layer links (Connecting across time/depth)
geom_segment(data = interlayer_segments, aes(x = x, y = y, xend = xend, yend = yend),
color = "steelblue", alpha = 0.4, size = 0.3, linetype = "dashed") +
# INTRA-layer edges
geom_edges(data = all_edges, aes(x = x, y = y, xend = xend, yend = yend),
color = "grey50", alpha = 0.2, size = 0.15) +
# Nodes
geom_nodes(data = all_edges, aes(x = x, y = y, color = layer), size = 1.3) +
scale_color_viridis_d(option = "plasma", end = 0.8) +
theme_void() +
coord_fixed(ratio = 0.8) + # Adjusted ratio to enhance the standing look
theme(legend.position = "none") +
annotate("text", x = offsets + 0.25, y = 1.6, label = layer_names, fontface = "bold")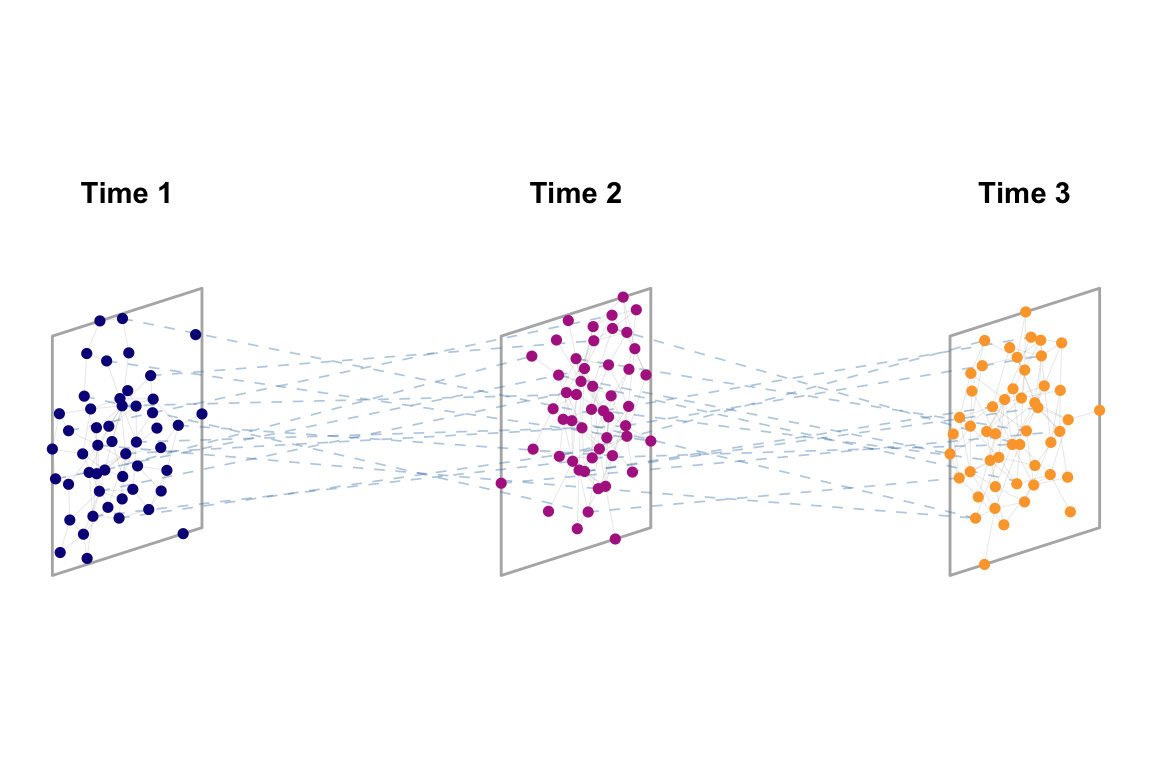
Plot the temporal network you chose to analyse in the last section.
4. Structural properties
MuxViz is largely the most comprehensive packlage fro multilayer network analysis. Here is a list of functions. It is impossible to go through all the structural properties available and you can see the references in the class page for papers and books.
Here is an example with in and out multi-degrees.
mEdges <- as.data.frame(multilayer_kefi$extended_ids[,c(2,1,4,3,5)])
colnames(mEdges)[1:4] <- c("node.from", "layer.from", "node.to", "layer.to")
M <- BuildSupraAdjacencyMatrixFromExtendedEdgelist(mEdges,3,106,T)
out_deg <- GetMultiOutDegree(M, Layers = 3, Nodes = 106, isDirected = T)
in_deg <- GetMultiInDegree(M, Layers = 3, Nodes = 106, isDirected = T)
tibble(out_deg, in_deg) %>% ggplot(aes(in_deg, out_deg))+geom_point()+labs(x='In degree', y='Out degree')+
geom_abline()+
xlim(0,85)+
ylim(0,85)+
coord_fixed()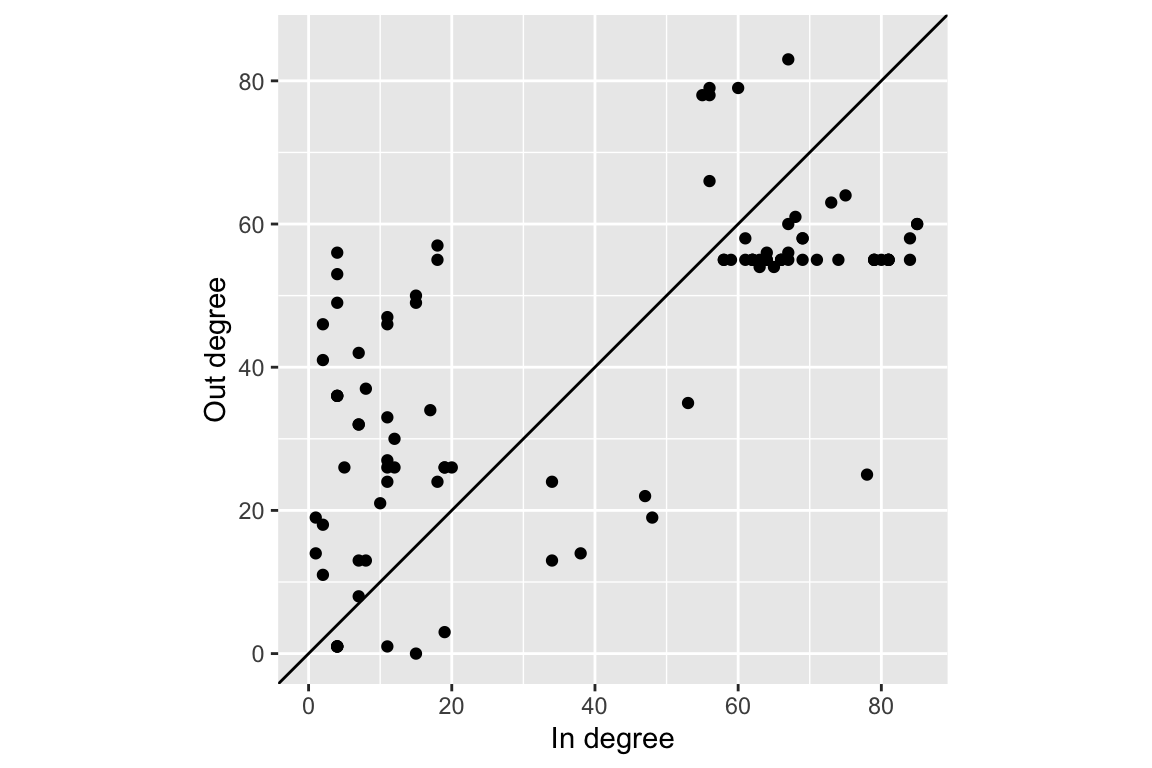
Try to program yourself two metrics from Boccaleti et al 2014:
- The total overlap \(O^{\alpha\beta}\) between two layers \(\alpha\) and \(\beta\), defined as the total number of links that are in common between the pair of layers.
- The local overlap \(o_i^{\alpha\beta}\) between two layers \(\alpha\) and \(\beta\), defined as the total number of neighbors of node \(i\) that are neighbors in both layers.
5. Aggregating networks
It is sometimes useful to aggregate a multilayer network into a single network. This is actually what has been traditionally done when not enough data were available. There are multiple ways to do so, as in the figure.
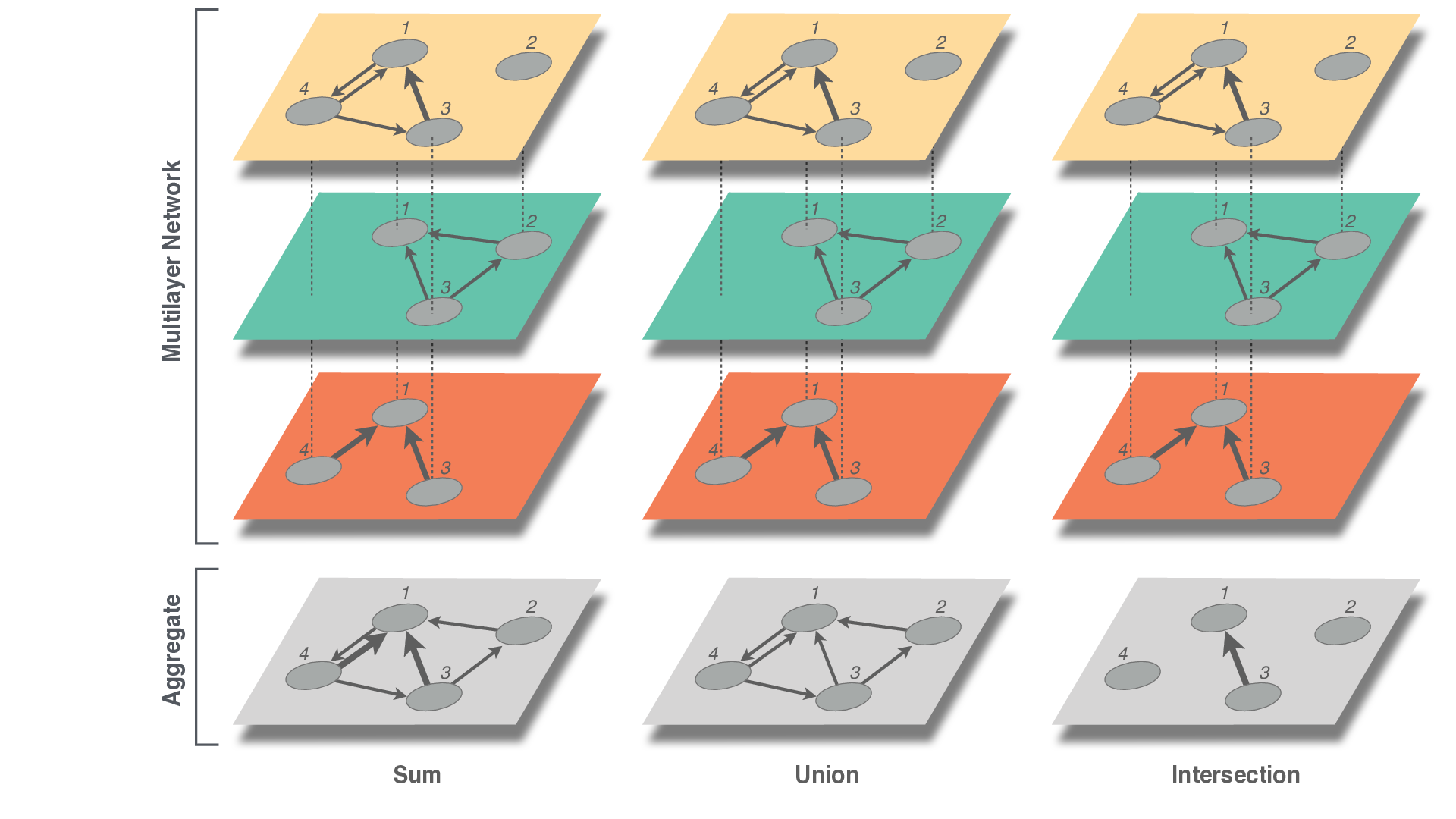
Write code to aggregate in each of these ways.
6. Community detection – Infomap
First, go over the example in the EMLN guide. This example does not have interlayer links so there is a global relax rate set.
Rerun the modularity while varying the global relax rate from 0.05 to 0.95 and record the number of modules. Plot the relationship. Is there a pattern? Can you explain it?

